- Log in to:
- Community
- DigitalOcean
- Sign up for:
- Community
- DigitalOcean
By Bharath K and Shaoni Mukherjee
Introduction
The applications of deep learning models and computer vision in the modern era are growing by leaps and bounds. Computer vision is one such field of artificial intelligence where we train our models to interpret real-life visual images. With the help of deep learning architectures like U-Net and CANet, we can achieve high-quality results on computer vision datasets to perform complex tasks. While computer vision is a humongous field with so much to offer and so many different, unique types of problems to solve, our focus for the next couple of articles will be on two architectures, namely U-Net and CANet, that are designed to solve the task of image segmentation.
Image segmentation involves dividing an image into multiple smaller segments or fragments, which simplifies and enhances the analysis of visual data. These segmented parts are essential for carrying out various image segmentation tasks. A key component in this process is the use of masks, binary images made up of zero and non-zero values, that help isolate specific regions of interest within an image. By applying these masks, we can accurately identify and work with the important elements in an image, paving the way for a wide range of downstream tasks.
Image segmentation has several vital applications, including machine vision, object detection, medical image analysis, and face recognition, among others. Before diving into the main content, we recommend reviewing a few optional prerequisites to better follow along. Below, you’ll find a table of contents outlining the key concepts covered in this article. While we encourage you to read through the entire guide, feel free to skip ahead to sections you’re already familiar with.
Prerequisites
- Python: Basic understanding of Python programming.
- Deep Learning: Familiarity with neural networks, particularly CNNs and object detection.
- PyTorch or TensorFlow: Knowledge of either framework is required to follow along.
Key Points
- Purpose of U-Net: Designed specifically for biomedical image segmentation, U-Net efficiently handles tasks where precise localization is required.
- Limitations of Traditional Methods: The sliding window approach introduced overlap and redundancy, making training slow and computationally expensive.
- Symmetric Architecture: U-Net features a symmetric encoder-decoder structure that captures context through downsampling and enables precise localization through upsampling.
- Skip Connections: These connections between encoder and decoder layers help retain spatial information lost during downsampling, improving segmentation accuracy.
- Mask Generation: The model produces segmentation masks, binary or multi-class, allowing for pixel-wise classification of input images.
- Versatility: Though initially developed for medical imaging, U-Net is widely used in domains such as satellite image analysis, industrial inspection, and autonomous driving.
- Data Efficiency: U-Net performs well even with limited training data, making it suitable for applications where annotated images are scarce.
Introduction To U-Net
The U-Net architecture, first published in 2015, has revolutionized deep learning. It won the International Symposium on Biomedical Imaging (ISBI) cell tracking challenge of 2015 in numerous categories by a large margin. Some of its works include the segmentation of neuron structures in electron microscopic stacks and transmitted light microscopy images.
With this U-Net architecture, the segmentation of images of size 512X512 can be computed with a modern GPU within a small amount of time. There have been many variants and modifications of this architecture due to its phenomenal success. Some of them include LadderNet, U-Net with attention, the recurrent and residual convolutional U-Net (R2-UNet), and U-Net with residual blocks or blocks with dense connections.
Although U-Net is a significant accomplishment in the field of deep learning, it is equally essential to understand the previous methods that were employed for solving such kind of similar tasks. One of the primary examples that comes to mind was the sliding window approach, which won the EM segmentation challenge at ISBI in the year 2012 by a large margin. The sliding window approach was able to generate a wide array of sample patches apart from the original training dataset.
This result was achieved by setting up the network of sliding window architecture by making the class label of each pixel a separate unit and providing a local region (patch) around that pixel. Another achievement of this architecture was that it could be localized quite easily on any given training dataset for the respective tasks.
However, the sliding window approach came with two significant limitations that were later addressed by the U-Net architecture. First, because each pixel was processed individually, the generated patches had substantial overlap, leading to a high degree of redundancy. Second, the training process was notably slow, requiring considerable time and computational resources. These challenges raised concerns about the overall efficiency and practicality of using such a network in real-world scenarios.
The U-Net is an elegant architecture that solves most of the issues. It uses the concept of fully convolutional networks for this approach. The intent of the U-Net is to capture both the features of the context as well as the localization. This process is completed successfully by the type of architecture built. The main idea of the implementation is to utilize successive contracting layers, which are immediately followed by the upsampling operators for achieving higher resolution outputs on the input images.
Understanding The U-Net Architecture
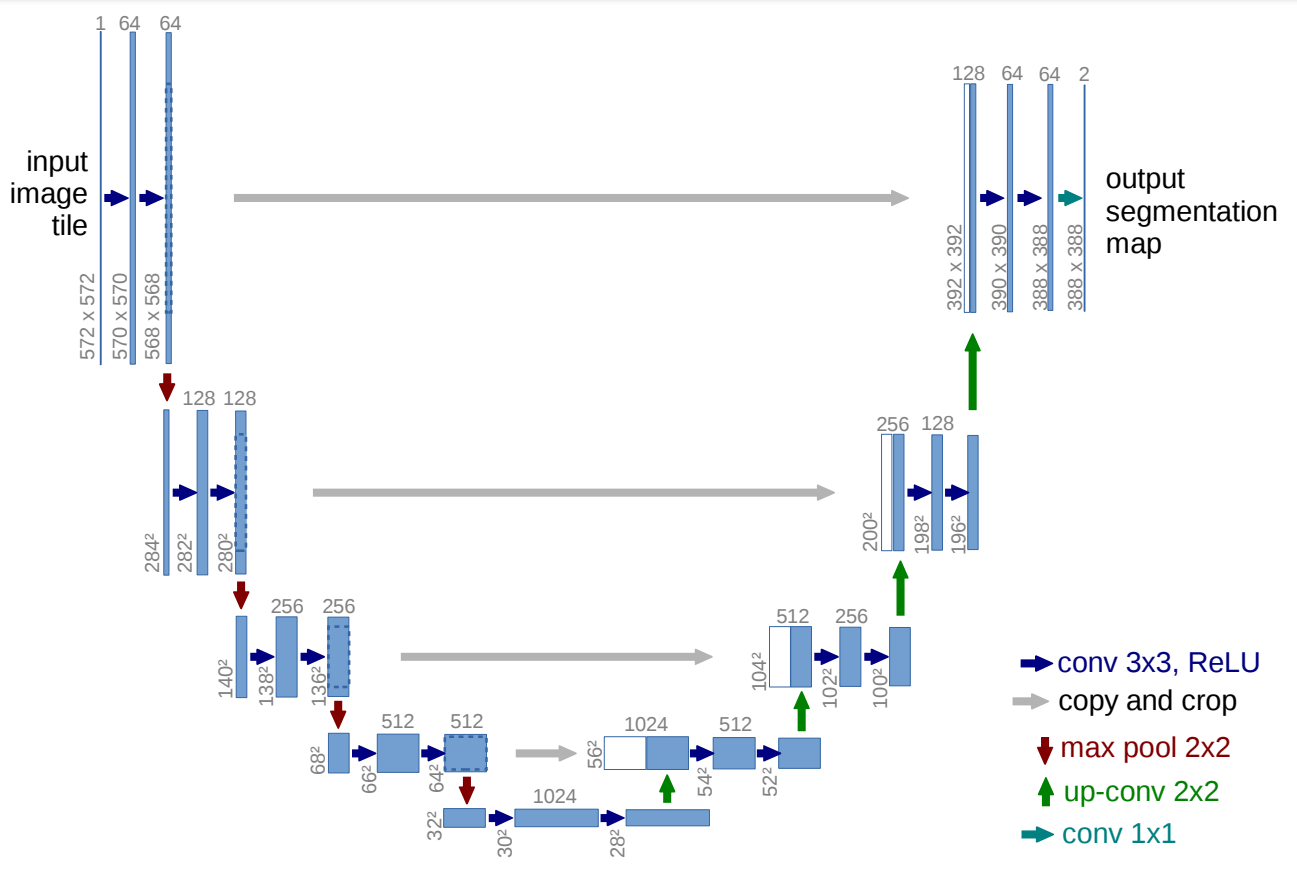
U-Net architecture from the paper “U-Net: Convolutional Networks for Biomedical Image Segmentation.”
By having a brief look at the architecture shown in the image, we can notice why it is probably referred to as U-Net architecture. The shape of the so-formed architecture is in the form of a 'U, and hence the following name. Just by looking at the structure and the numerous elements involved in the process of the construction of this architecture, we can understand that the network built is a fully convolutional network. They have not used any other layers, such as dense or flattened, or other similar layers. The visual representation shows an initial contracting path followed by an expanding path.
The architecture shows that an input image is passed through the model, and then it is followed by a couple of convolutional layers with the ReLU activation function. We can notice that the image size is reducing from 572X572 to 570X570 and finally to 568X568. The reason for this reduction is that they have made use of unpadded convolutions (defined the convolutions as “valid”), which results in the reduction of the overall dimensionality. Apart from the Convolution blocks, we also notice that we have an encoder block on the left side, followed by a decoder block on the right side.
The encoder block has a constant reduction of image size with the help of the max-pooling layers of strides 2. We also have repeated convolutional layers with an increasing number of filters in the encoder architecture. Once we reach the decoder aspect, we notice the number of filters in the convolutional layers starts to decrease along with a gradual upsampling in the following layers, all the way to the top. We also notice that the use of skip connections connects the previous outputs with the layers in the decoder blocks.
This skip connection is vital to preserve the loss from the previous layers so that they reflect stronger on the overall values. It is also scientifically proven to produce better results and lead to faster model convergence. In the final convolution block, we have a couple of convolutional layers followed by the final convolution layer. This layer has a filter of 2 with the appropriate function to display the resulting output. This final layer can be changed according to the desired purpose of the project you are trying to perform.
TensorFlow Implementation of U-Net
In this section of the article, we will look at the TensorFlow implementation of the U-Net architecture. While I am utilizing TensorFlow for the computation of the model, you can choose any deep learning framework, such as PyTorch, for a similar implementation. We will look at the working of the U-Net architecture along with some other model structures with PyTorch in future articles. However, for this article, we will stick to the TensorFlow library. We will be importing all the required libraries and constructing our U-Net architecture from scratch. But we will make some necessary changes that will improve the overall performance of the model as well as make it slightly less complex.
Modifications in the implemented model
It is crucial to note that the U-Net model was introduced way back in 2015. Although its performance at that point in time was fabulous, the prominent methods and functions of deep learning have evolved simultaneously as well. Hence, there have been many successful variants and versions of the U-Net architecture since its original creation to preserve certain image qualities while replicating, and in some scenarios, performing better than the original architecture.
I will try to preserve most of the essential parameters and architectural elements of the original implementation of the U-Net architecture. However, there will be slight changes from the original content that will improve the modern efficiency and speed, as well as the simplicity of the model. One of the changes that will be included in this structure is using the value of Convolution as “same” because many pieces of research in the future have shown that this particular change did not negatively impact the architectural build in any way. Also, since the concept of batch normalization was introduced in 2016, the original architecture did not use this aspect. But our model implementation will include batch normalization as it yields the best results in most cases.
Importing the required libraries
For building the U-Net architecture, we will utilize the TensorFlow deep learning framework, as discussed already. Hence, we will import the TensorFlow library for this purpose as well as the Keras framework, which is now an integral part of TensorFlow model structures. From our previous understanding of the U-Net architecture, we know that some of the essential imports include the convolutional layer, the max-pooling layer, an input layer, and the activation function ReLU for the basic modeling structure. We will then use some additional layers like the Conv2DTranspose layer, which will perform an upsampling for our desired decoder blocks. We will also make use of the Batch Normalization layers for stabilizing the training process and use the Concatenate layers for combining the necessary skip connections.
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.layers import Input, Conv2D, MaxPooling2D, Activation, ReLU
from tensorflow.keras.layers import BatchNormalization, Conv2DTranspose, Concatenate
from tensorflow.keras.models import Model, Sequential
Building the Convolution Block
After importing the required libraries, we can continue to build the U-Net architecture. You can either do this in one complete class by defining all the parameters and values accordingly in order and continuing the process until you reach the very end, or you a few iterative blocks. I will be using the latter method as it is more convenient for most users to understand the model architecture of U-Net with the help of a few blocks. We will utilize three iterative blocks as shown in the architecture representation, namely the convolution operation block, the encoder block, and the decoder block. With the help of these three blocks, we can build the U-Net architecture with ease. Let us now process and understand each of these function code blocks one by one.
The convolution operation block is used to perform the primary operation of taking the entered input parameters and processing a double layer of convolution operations. In this function, we have two arguments, namely the input for the convolution layer and the number of filters, which is by default 64. We will use the value of padding as same as discussed previously to maintain the same shapes as opposed to unpadded or valid convolutions. These convolutional layers are followed by the Batch Normalization layer. These changes from the original model are made to gain the best outcomes possible. Finally, a ReLU activation layer is added to the mix as defined in the research paper. Let us explore the code block for building the convolution block.
def convolution_operation(entered_input, filters=64):
# Taking first input and implementing the first conv block
conv1 = Conv2D(filters, kernel_size = (3,3), padding = "same")(entered_input)
batch_norm1 = BatchNormalization()(conv1)
act1 = ReLU()(batch_norm1)
# Taking first input and implementing the second conv block
conv2 = Conv2D(filters, kernel_size = (3,3), padding = "same")(act1)
batch_norm2 = BatchNormalization()(conv2)
act2 = ReLU()(batch_norm2)
return act2
Constructing the encoder and decoder blocks
Our Next step will be to build the encoder and decoder blocks. These two functions are quite simple to construct. The encoder architecture will use consecutive inputs starting from the first layer all the way to the bottom. The encoder function as we have defined will have the convolutional block, i.e., two convolutional layers followed by their respective batch normalization and ReLU layers. Once we pass them through the convolution blocks, we will quickly downsample these elements, as mentioned in the research paper. We will use a max-pooling layer and stick to the parameters mentioned in the paper with strides = 2. We will then return both the initial output and the max-pooled output, as we need the former for performing the skip connections.
The decoder block will include three arguments, namely the receiving inputs, the input of the skip connection, and the number of filters in the particular building block. We will upsample the entered input with the help of the Conv2DTranspose layers in our model. We will then concatenate both the receiving input and the newly upsampled layers to receive the final value of the skip connections. We will then use this combined function and perform our convolutional block operation to proceed to the next layer and return this output value.
def encoder(entered_input, filters=64):
# Collect the start and end of each sub-block for normal pass and skip connections
enc1 = convolution_operation(entered_input, filters)
MaxPool1 = MaxPooling2D(strides = (2,2))(enc1)
return enc1, MaxPool1
def decoder(entered_input, skip, filters=64):
# Upsampling and concatenating the essential features
Upsample = Conv2DTranspose(filters, (2, 2), strides=2, padding="same")(entered_input)
Connect_Skip = Concatenate()([Upsample, skip])
out = convolution_operation(Connect_Skip, filters)
return out
Construct the U-Net architecture
If you are trying to build the entire U-Net architecture from scratch in a single layer, you might find that the overall structure is quite humongous because it consists of so many different blocks to be processed. By dividing our respective functions into three separate code blocks of convolutional operation, encoder structure, and decoder structure, we can construct the U-Net architecture with ease in a few lines of code. We will use the input layer, which will contain the respective shapes of our input image.
After this step, we will collect all the primary outputs and the skip outputs to pass them on to further blocks. We will create the next block and construct the entire decoder architecture until we reach the output. The output will have the required dimensions according to our desired output. In this case, I have one output node with the sigmoid activation function. We will call the functional API modeling system to create our final model and return this model to the user for performing any task with the U-Net architecture.
def U_Net(Image_Size):
# Take the image size and shape
input1 = Input(Image_Size)
# Construct the encoder blocks
skip1, encoder_1 = encoder(input1, 64)
skip2, encoder_2 = encoder(encoder_1, 64*2)
skip3, encoder_3 = encoder(encoder_2, 64*4)
skip4, encoder_4 = encoder(encoder_3, 64*8)
# Preparing the next block
conv_block = convolution_operation(encoder_4, 64*16)
# Construct the decoder blocks
decoder_1 = decoder(conv_block, skip4, 64*8)
decoder_2 = decoder(decoder_1, skip3, 64*4)
decoder_3 = decoder(decoder_2, skip2, 64*2)
decoder_4 = decoder(decoder_3, skip1, 64)
out = Conv2D(1, 1, padding="same", activation="sigmoid")(decoder_4)
model = Model(input1, out)
return model
Finalizing the Model
Ensure that your image shapes are divisible by at least 16 or multiples of 16. Since we are using four max-pooling layers during the down-sampling procedure, we don’t want to encounter the divisibility of any odd-numbered shapes. Hence, it would be best to ensure that the sizes of your architecture are equivalent to sizes like (48, 48), (80,80), (160, 160), (256, 256), (512, 512), and other similar shapes. Let us try our model structure for an input shape of (160, 160, 3) and test the results.
input_shape = (160, 160, 3)
model = U_Net(input_shape)
model.summary()
tf.keras.utils.plot_model(model, "model.png", show_shapes=False, show_dtype=False, show_layer_names=True, rankdir='TB', expand_nested=False, dpi=96)
You can view the summary and plots respectively with the above code blocks. Let us now explore a fun project with the U-Net architecture.
Quick Example Project To View U-Net Performance
For this project, we will use the reference from Keras for an image segmentation project. For this project, we will extract the dataset and visualize the basic elements to get an overview of the structure. We can then proceed to build the data generator for loading data from the dataset accordingly. We will then utilize the U-Net model that we built in the previous section and train this model until we reach a satisfactory result. Once our desired result is obtained, we will save this model and test it out on a validation sample. Let us get started with the implementation of the project!
Dataset Preparation
We will use the Oxford pet dataset for this particular task of image segmentation. The Oxford pet dataset consists of 37 categories of pets with roughly 200 images for each class. The images have a large variation in scale, pose, and lighting. Once you have the zip files downloaded successfully, you can unzip them twice with 7-Zip or any other similar tool kit that you use on your operating system.
In the first code block, we will define the respective paths to the images and annotations directories. We will also define some essential parameters such as the image size, the batch size, and the number of classes. We will then ensure that all the elements in the dataset are arranged in the correct order for performing and obtaining a satisfactory image segmentation task. You can verify the images with their respective annotations by printing both the file paths to check if they produce the desired results.
import os
input_dir = "images/"
target_dir = "annotations/trimaps/"
img_size = (160, 160)
num_classes = 3
batch_size = 8
input_img_paths = sorted(
[
os.path.join(input_dir, fname)
for fname in os.listdir(input_dir)
if fname.endswith(".jpg")
]
)
target_img_paths = sorted(
[
os.path.join(target_dir, fname)
for fname in os.listdir(target_dir)
if fname.endswith(".png") and not fname.startswith(".")
]
)
print("Number of samples:", len(input_img_paths))
for input_path, target_path in zip(input_img_paths[:10], target_img_paths[:10]):
print(input_path, "|", target_path)
Data Visualization
Now that we have collected and pre-processed our data for our project, our next step will be to take a brief look at the dataset. Let us analyze the dataset by displaying both an image and its respective segmented output. This segmented output with the masking is often referred to as the ground truth annotation. We will utilize the I-Python display option along with the Pillow library for randomly displaying a selected image. This simple code block is written as follows:
from IPython.display import Image, display
from tensorflow.keras.preprocessing.image import load_img
import PIL
from PIL import ImageOps
# Display input image #7
display(Image(filename=input_img_paths[9]))
# Display auto-contrast version of corresponding target (per-pixel categories)
img = PIL.ImageOps.autocontrast(load_img(target_img_paths[9]))
display(img)


Configure the data generator
In this particular code block, instead of using data generators to prepare the data, we will utilize the Sequence operation from the Keras deep learning framework. This method is much safer for multi-processing in comparison to using the generator because it only trains each sample once per epoch, which will avoid any unnecessary adjustments that we will need to make. We will then prepare the Sequence class in the utils module of Keras to compute, load, and vectorize batches of data. We will then construct the initialization function, an additional function to compute the length, and the final function that will generate batches of data.
from tensorflow import keras
import numpy as np
from tensorflow.keras.preprocessing.image import load_img
class OxfordPets(keras.utils.Sequence):
"""Helper to iterate over the data (as Numpy arrays)."""
def __init__(self, batch_size, img_size, input_img_paths, target_img_paths):
self.batch_size = batch_size
self.img_size = img_size
self.input_img_paths = input_img_paths
self.target_img_paths = target_img_paths
def __len__(self):
return len(self.target_img_paths) // self.batch_size
def __getitem__(self, idx):
"""Returns tuple (input, target) correspond to batch #idx."""
i = idx * self.batch_size
batch_input_img_paths = self.input_img_paths[i : i + self.batch_size]
batch_target_img_paths = self.target_img_paths[i : i + self.batch_size]
x = np.zeros((self.batch_size,) + self.img_size + (3,), dtype="float32")
for j, path in enumerate(batch_input_img_paths):
img = load_img(path, target_size=self.img_size)
x[j] = img
y = np.zeros((self.batch_size,) + self.img_size + (1,), dtype="uint8")
for j, path in enumerate(batch_target_img_paths):
img = load_img(path, target_size=self.img_size, color_mode="grayscale")
y[j] = np.expand_dims(img, 2)
# Ground truth labels are 1, 2, 3. Subtract one to make them 0, 1, 2:
y[j] -= 1
return x, y
In the next step, we will define the split between the training and the validation data, respectively. We do this step to ensure that there is no corruption between the integrity of the elements in the train and test sets. Both of these data entities must be viewed separately so that the model does not get a peek at the testing data. In the validation set, we will also perform an optional shuffle operation that will mix up all the images in the dataset, and we can obtain random samples for both the training and the validation images. We will then call the training values and the validation values individually and store them in their respective variables.
import random
# Split our img paths into a training and a validation set
val_samples = 1000
random.Random(1337).shuffle(input_img_paths)
random.Random(1337).shuffle(target_img_paths)
train_input_img_paths = input_img_paths[:-val_samples]
train_target_img_paths = target_img_paths[:-val_samples]
val_input_img_paths = input_img_paths[-val_samples:]
val_target_img_paths = target_img_paths[-val_samples:]
# Instantiate data Sequences for each split
train_gen = OxfordPets(
batch_size, img_size, train_input_img_paths, train_target_img_paths
)
val_gen = OxfordPets(batch_size, img_size, val_input_img_paths, val_target_img_paths)
Once we have completed the following steps, we can proceed to construct our U-Net architecture.
U-Net Model
The U-Net model we will construct in this section is the exact same architecture as the one defined in the previous sections, except for a few small modifications that we will discuss shortly. After the preparation of the dataset, we can construct our model accordingly. The model inculcates the image sizes and starts to structure the overall architecture through which our images will be passed. The only change you will need to make in the previously discussed architecture is as follows:
out = Conv2D(3, 1, padding="same", activation="sigmoid")(decoder_4)
Or
out = Conv2D(num_classes, 1, padding="same", activation="sigmoid")(decoder_4)
We are changing the final layer in the U-Net architecture to signify the total number of outputs that will be generated at the final step. Note that you could also utilize the SoftMax function for generating the final output for multi-class classification, and that is probably more accurate. However, you can see from the training results that this activation function works fine as well.
Train the model
In the next step, we will compile and train the model to see its performance on the data. We are also using a checkpoint to save the model so that we can make predictions in the future. I interrupted the training procedure after 11 epochs as I was quite satisfied with the result obtained. You can choose to run it for more epochs if you want.
model.compile(optimizer="rmsprop", loss="sparse_categorical_crossentropy")
callbacks = [
keras.callbacks.ModelCheckpoint("oxford_segmentation.h5", save_best_only=True)
]
# Train the model, doing validation at the end of each epoch.
epochs = 15
model.fit(train_gen, epochs=epochs, validation_data=val_gen, callbacks=callbacks)
Epoch 1/15
798/798 [==============================] - 874s 1s/step - loss: 0.7482 - val_loss: 0.7945
Epoch 2/15
798/798 [==============================] - 771s 963ms/step - loss: 0.4964 - val_loss: 0.5646
Epoch 3/15
798/798 [==============================] - 776s 969ms/step - loss: 0.4039 - val_loss: 0.3900
Epoch 4/15
798/798 [==============================] - 776s 969ms/step - loss: 0.3582 - val_loss: 0.3574
Epoch 5/15
798/798 [==============================] - 788s 985ms/step - loss: 0.3335 - val_loss: 0.3607
Epoch 6/15
798/798 [==============================] - 778s 972ms/step - loss: 0.3078 - val_loss: 0.3916
Epoch 7/15
798/798 [==============================] - 780s 974ms/step - loss: 0.2772 - val_loss: 0.3226
Epoch 8/15
798/798 [==============================] - 796s 994ms/step - loss: 0.2651 - val_loss: 0.3046
Epoch 9/15
798/798 [==============================] - 802s 1s/step - loss: 0.2487 - val_loss: 0.2996
Epoch 10/15
798/798 [==============================] - 807s 1s/step - loss: 0.2335 - val_loss: 0.3020
Epoch 11/15
798/798 [==============================] - 797s 995ms/step - loss: 0.2220 - val_loss: 0.2801
View the results
Finally, let us visualize the results obtained.
# Generate predictions for all images in the validation set
val_gen = OxfordPets(batch_size, img_size, val_input_img_paths, val_target_img_paths)
val_preds = model.predict(val_gen)
def display_mask(i):
"""Quick utility to display a model's prediction."""
mask = np.argmax(val_preds[i], axis=-1)
mask = np.expand_dims(mask, axis=-1)
img = PIL.ImageOps.autocontrast(keras.preprocessing.image.array_to_img(mask))
display(img)
# Display results for validation image #10
i = 10
# Display input image
display(Image(filename=val_input_img_paths[i]))
# Display ground-truth target mask
img = PIL.ImageOps.autocontrast(load_img(val_target_img_paths[i]))
display(img)
# Display mask predicted by our model
display_mask(i) # Note that the model only sees inputs at 150x150.



FAQ’s
Q: U-Net vs DeepLab vs Mask R-CNN: which segmentation model to choose in 2025?
A: Choose segmentation models based on specific requirements:
- U-Net excels for biomedical and scientific imaging with limited data, provides excellent boundary preservation through skip connections, and works well with small datasets.
- DeepLab offers superior performance on natural images, handles multiple scales effectively through atrous convolutions, and provides good efficiency for real-time applications.
- Mask R-CNN combines object detection with instance segmentation, ideal for applications requiring both object identification and precise boundaries.
- Performance: DeepLab typically achieves highest accuracy on natural images, U-Net dominates medical imaging, Mask R-CNN best for instance segmentation.
- Data requirements: U-Net works with 100-500 images, DeepLab needs 1000+, Mask R-CNN requires extensive labeled data.
- Computational needs: U-Net most efficient, DeepLab moderate, Mask R-CNN most demanding.
Q: How to implement attention mechanisms in U-Net for better segmentation accuracy?
A: Attention mechanisms enhance U-Net performance through focused feature processing:
- Attention U-Net adds attention gates between encoder-decoder skip connections to suppress irrelevant features and highlight important regions.
- Self-attention modules can be integrated into encoder or decoder paths for global context modeling.
- Channel attention (SE blocks) improves feature representation by learning channel-wise importance.
- Spatial attention focuses on relevant spatial locations within feature maps.
- Implementation: Add attention gates before skip connections using sigmoid gating mechanisms.
- Multi-scale attention processes features at different resolutions for comprehensive context.
- Transformer integration using vision transformer blocks within U-Net architecture.
- Benefits: 2-5% improvement in IoU scores, better handling of complex scenes, improved boundary precision.
- Code example: Implement attention gates as separate modules that take both encoder and decoder features as input.
Q: What data augmentation techniques work best for medical image segmentation with U-Net?
A: Medical image segmentation requires specialized augmentation strategies:
- Geometric transforms: Random rotation (±15°), elastic deformation for anatomical variation, scaling and translation within reasonable bounds.
- Intensity augmentation: Gaussian noise addition, contrast/brightness adjustment, gamma correction for different imaging conditions.
- Spatial augmentation: Random cropping with proper mask alignment, flipping (horizontal/vertical as appropriate for anatomy).
- Medical-specific: Simulated motion artifacts, imaging noise patterns, contrast agent variations.
- Advanced techniques: Mixup for segmentation, random erasing of image regions, synthetic data generation using GANs.
- Implementation considerations: Ensure augmentations maintain anatomical realism, apply same transforms to image and mask, validate augmentations don’t introduce artifacts. * Libraries: Albumentations, TorchIO for medical-specific augmentations. Balance: Strong augmentation prevents overfitting but excessive augmentation can harm performance.
Q: How to optimize U-Net training for large high-resolution medical images?
A: Training U-Net on large medical images requires memory and computational optimizations:
- Patch-based training dividing large images into overlapping patches (256x256 or 512x512) for manageable memory usage.
- Progressive training starting with low resolution and gradually increasing input size.
- Mixed precision training using FP16 to reduce memory consumption by ~50%.
- Gradient accumulation for effective large batch training on limited GPU memory.
- Efficient data loading with prefetching, multiple workers, and memory mapping for large datasets.
- Model optimizations: Depthwise separable convolutions, channel reduction in skip connections, efficient attention mechanisms.
- Multi-GPU training using data parallelism for faster training.
- Memory management: Clear unnecessary variables, use checkpoint-based backpropagation, implement efficient data structures.
- Inference optimization: Sliding window prediction with overlap handling for seamless reconstruction.
Q: What are the latest improvements to U-Net architecture for segmentation tasks?
A: Modern U-Net variants incorporate several architectural advances:
- U-Net++ uses dense skip connections and deep supervision for better feature fusion and gradient flow.
- Attention U-Net integrates attention mechanisms for improved focus on relevant features.
- U-Net 3+ implements full-scale skip connections connecting all encoder levels to all decoder levels.
- TransUNet combines CNN encoders with transformer decoders for global context modeling.
- nnU-Net provides automatic architecture optimization and hyperparameter tuning for medical segmentation.
- Swin-UNet uses Swin Transformer blocks for both encoder and decoder paths.
- Performance improvements: These variants typically achieve 2-8% higher IoU scores than standard U-Net.
- Implementation: Most improvements focus on better skip connections, attention mechanisms, and multi-scale feature processing.
- Considerations: Advanced variants require more computational resources and careful hyperparameter tuning but provide superior segmentation quality.
Conclusion
The U-Net architecture is one of the most significant and revolutionary landmarks in the field of deep learning. While the initial research paper that introduced the U-Net architecture was to solve the task of Biomedical Image Segmentation, it was not limited to this single application. The model could and can still solve the most complex problems in deep learning. Although some of the elements in the original architecture are outdated, there are several variations of this architecture. These include LadderNet, U-Net with attention, the recurrent and residual convolutional U-Net (R2-UNet), and other similar networks, which are derived successfully from the original U-Net Models.
In this article, we looked into a brief introduction of the U-Net modeling technique that finds fabulous utility in most modern tasks related to image segmentation. We then proceed to understand the construction and the main methodologies employed in the building of the U-Net architecture. We understood the numerous methodologies and techniques used to achieve the best results possible on the provided dataset. After this section, we learned how to build the U-Net architecture from scratch with various blocks to simplify the overall structure. Finally, we analyzed our constructed U-Net architecture with a simple example problem for image segmentation.
Thanks for learning with the DigitalOcean Community. Check out our offerings for compute, storage, networking, and managed databases.
About the author(s)
With a strong background in data science and over six years of experience, I am passionate about creating in-depth content on technologies. Currently focused on AI, machine learning, and GPU computing, working on topics ranging from deep learning frameworks to optimizing GPU-based workloads.
Still looking for an answer?
This textbox defaults to using Markdown to format your answer.
You can type !ref in this text area to quickly search our full set of tutorials, documentation & marketplace offerings and insert the link!
- Table of contents
- Introduction
- Prerequisites
- Introduction To U-Net
- Understanding The U-Net Architecture
- TensorFlow Implementation of U-Net
- Construct the U-Net architecture
- Quick Example Project To View U-Net Performance
- Train the model
- FAQ's
- Conclusion
Limited Time: Introductory GPU Droplet pricing.
Get simple AI infrastructure starting at $2.99/GPU/hr on-demand. Try GPU Droplets now!
Become a contributor for community
Get paid to write technical tutorials and select a tech-focused charity to receive a matching donation.
DigitalOcean Documentation
Full documentation for every DigitalOcean product.
Resources for startups and AI-native businesses
The Wave has everything you need to know about building a business, from raising funding to marketing your product.
Get our newsletter
Stay up to date by signing up for DigitalOcean’s Infrastructure as a Newsletter.
New accounts only. By submitting your email you agree to our Privacy Policy
The developer cloud
Scale up as you grow — whether you're running one virtual machine or ten thousand.
Start building today
From GPU-powered inference and Kubernetes to managed databases and storage, get everything you need to build, scale, and deploy intelligent applications.
